Molecular and Developmental Biology
Faculty research interests in the area of molecular and developmental biology include evolutionary dervelopmental biology of vertebrates (Diaz Lab), developmental biology and cell signaling (Eivers Lab), molecular mechanisms in vertebrate development (Nissen Lab), and molecular regulatory mechanisms of development (Sharp Lab). An asterisk following a name in a publication citation indicates a student coauthor.
Evolutionary Developmental Biology of Vertebrates
| Contact: Raul Diaz, Ph.D. Office: ASCL 394, ext: 3-2077 Laboratory: ASCL 309 E-mail: [email protected] |
||
|
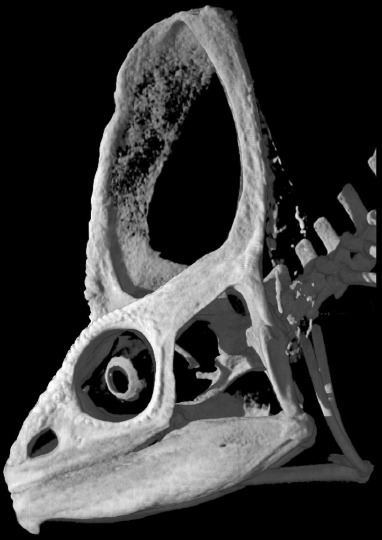
This image (data from digimorph.org) shows the unique skull anatomy present across chameleons where bones grow vertically away from the skull roof much like a tent, a unique architecture only seen in the extinct lineage of ceratopsid dinosaurs. Our lab aims to study how such unique morphologies develop and evolve and also examine how living organisms can provide us insight into better understanding species that have gone extinct. |
Research Summary
My lab is interested in studying the diversity of body plans present across vertebrates (morphological disparity), but more specifically we are interested in the development and diversification of reptiles. Our sampling spans embryos produced in house as well as fieldwork both within and outside the USA. In addition, we employ a very broad array of classic and modern lab and genomic techniques to understand embryonic morphogenesis from a genetic to cellular and tissue level while also understanding how organisms interact with their environments by placing structural anatomy within the framework of biomechanics, ecology and phylogeny. Insights into the evolution of body plans can also come from understanding human diseases and lab based disease models, where anatomical changes associated with malformations are disadvantageous (or lethal) in some organisms, but are adaptive in other organisms and ecological contexts such as the cleft/split hands/feet of chameleons. This latter research area we have called Evo Devo Path and is also the study of “Hopeful Monsters.”
| Representative Publications |
|
R. E. Diaz, Jr, N. A. Shylo, D. Roellig, M. Bronner and P. A. Trainor. 2019. Filling in the phylogenetic gaps: induction, migration and differentiation of neural crest cells in a squamates reptile, the veiled chameleon (Chamaeleo calyptratus). Developmental Dynamics |
|
Pinto, B. J., D. C. Card, T. A. Castoe, R. E. Diaz, Jr., S. V. Nielsen, P. A. Trainor and T. Gamble. 2019. The transcriptome of the veiled chameleon (Chamaeleo calyptratus): a resource for studying the evolution and development of vertebrates. Developmental Dynamics. |
|
Diaz, Jr., R. E. and P. A. Trainor. 2018. An integrative view of Lepidosaur cranial anatomy, development and diversification in Heads, Jaws, and Muscles: Anatomical, Functional, and Developmental Diversity in Chordate Evolution, Chapter 9. Eds: Janine M. Ziermann, Raul E. Diaz, Jr., Rui Diogo. Springer Publishers: Fascinating Life Sciences Series. |
|
Diaz, Jr., R. E., F. Bertocchini, and P. Trainor. 2017. Lifting the veil on reptile embryology: the veiled chameleon (Chamaeleo calyptratus) as a model system to study reptilian development. In Avian and Reptilian Developmental Biology (Methods in Molecular Biology, Volume 1650), pp. 269-284. |
|
Molnar, J. L., R. E. Diaz, Jr., T. Skorka, G. Dagliyan, and R. Diogo. 2017. Comparative musculoskeletal anatomy of chameleon limbs, with implications for the evolution of arboreal locomotion in lizards and for teratology. Journal of Morphology |
Developmental Biology and Cell Signaling
 |
Contact: Edward Eivers, Ph.D. Office: ASCL 392, ext: 3-2075 Laboratory: ASCL 350, ext: 3-2071 E-mail: [email protected] |
|
| Fluorescent immunostaining of phosphorylated Mad in the dorsal most cells of a Drosophila blastoderm embryo. Phosphorylated Mad represents the cells which receive high levels of BMP signals. BMP target genes are strongly activated in this region. |
Research Summary
My research lab investigates signaling crosstalk between the bone morphogenetic protein (BMP) and Wnt signaling pathways during Drosophila and Xenopus embryonic development. Both signaling pathways are evolutionarily conserved from flies to mammals and regulate a diverse range of biological events. These include stem cell maintenance, cell differentiation and organogenesis, while abnormal signaling of either pathway has strongly been linked to various forms of cancer. I am particularly interested in how the transcription factor Mad, which is required for both BMP and Wnt signaling, determines cell fate and differentiation during early tissue development.
| Representative Publications |
|
Urrutia, H., Aleman, A. and Eivers E. 2016. Drosophila Dullard functions as a Mad phosphatase to terminate BMP signaling. Sci. Rep. 6, 32269; doi: 10.1038/srep32269 |
|
Aleman, A., Rios, M., Juarez, M., Lee, D., Chen, A. and Eivers, E. 2014. Mad linker phosphorylations control the intensity and range of the BMP-activity gradient in developing Drosophila tissues. Scientific Reports 4: 6927; DOI:10.1038/srep06927. |
|
Eivers, E., Demagny, H., Choi, R.H., and De Robertis, E.M. 2011. Phosphorylation of Mad controls competition between Wingless and BMP signaling. Science Signaling 4: ra68. |
| Fuentealba, L.C., Eivers, E., Geissert, D., Taelman, V., and De Robertis, E.M. 2008. Asymmetric mitosis: Unequal segregation of proteins destined for degradation. Proceedings of the Nationall Academy of Sciences USA 105: 7732-7737. |
| Fuentealba, L.C., Eivers, E., Ikeda, A., Hurtado, C., Kuroda, H., Pera, E.M., and De Robertis, E.M. 2007. Integrating patterning signals: Wnt/GSK3 regulates the duration of the BMP/Smad1 signal. Cell 131: 980-993. |
Molecular Mechanisms in Vertebrate Development
 |
Contact: Robert Nissen, Ph.D. Office: ASCL 216, ext: 3-2309 Laboratory: ASCL 210, ext: 3-2093 E-mail: [email protected] Web: /faculty/rnissen |
|
| Example of a wild type zebrafish embryo at 28 hours. Zebrafish are used as a model organism for studying the molecular mechanisms of vertebrate development. |
Research Summary
Our long-term goal is to understand the mechanisms by which an initially pluripotent cell becomes restricted to specific fates and subsequently how it is maintained in the differentiated state. Currently, our focus is aimed at revealing the roles that the Wdr68 and Dyrk1b proteins play in the Nodal signaling pathway using zebrafish as a model organism. Ultimately, we hope these studies will shed new and valuable light on the mechanisms driving cell specification and differentiation in vertebrates.
| Representative Publications |
| Mazmanian, G.*, Kovshilovsky, M.*, Yen, D.*, Mohanty, A.*, Mohanty, S.*, Nee, A., and Nissen, R.M. 2010. The zebrafish dyrk1b gene is important for endoderm formation. Genesis 48: 20. |
| Nissen, R.M., Amsterdam, A., and Hopkins, N. 2006. A zebrafish screen for craniofacial mutants identifies wdr68 as a highly conserved gene required for endothelin-1 expression. BMC Developmental Biology 6: 28. |
| Mansfield, J.H., Harfe, B.D., Nissen, R.M., Obenauer, J., Srineel, J., Chaudhuri, A., Farzan-Kashani, R., Zucker, M., Pasquinelli, A.E., Ruvkin, G., Sharp, P.A., Tabin, C.J., and McManus, M.T. 2004. MicroRNA-responsive "sensor" transgenes reveal Hox-like and other developmentally regulated patterns of vertebrate microRNA expression. Nature Genetics 36: 1079-1083. |
| Amsterdam, A., Nissen, R.M., Sun, Z., Swindell, E.C., Farrington, S., and Hopkins, N. 2004. 315 genes essential for early zebrafish development. Proceedings of the National Academy of Sciences 101: 12792-12797. |
| Nissen, R.M., Yan, J., Amsterdam, A., Hopkins, N., and Burgess, S. 2003. Zebrafish foxi one modulates cellular responses to FGF signaling required for the integrity of ear and jaw patterning. Development 130: 2543-2554. |
Molecular Regulatory Mechanisms of Development
 |
Contact: Sandra Sharp, Ph.D. Office: ASCL 316, ext: 3-2072 Laboratory: ASCL 328, ext: 3-4820 E-mail: [email protected] Web: /faculty/ssharp |
|
| AlexaFluor immuno-stained muscle fibers in a section of normal mouse hindlimb. Both cross and nearly longitudinal views are visible. |
Research Summary
An important goal in biomedical research is to understand the molecular pathways and mechanisms that regulate development, particularly those that regulate the transition from cell proliferation to cell differentiation. Our model system is muscle development or myogenesis. We currently have two major projects supported by grant funding from the National Institutes of Health. The first uses whole animals and molecular studies to test the hypothesis that either the generally expressed tumor suppressor transcription factor p53 or the muscle-specific transcription factor MyoD must be expressed for successful completion of myogenesis in vivo. The second uses an ultra-high throughput ChIP-Seq approach to understand the regulatory mechanisms involved in controlling binding of MyoD to DNA as cells stop dividing and develop into muscle. We aspire to contribute to the understanding of the mechanisms that regulate normal differentiation in order to enable the development of successful medical approaches to both developmental disorders and cancer.
| Representative Publications |
| Krilowicz, B., Johnston, W., Sharp, S.B., Warter-Perez, N., Momand, J. 2007. A summer program designed to educate college students for careers in bioinformatics. CBE-Life Sciences Education 6: 74-83. |
| Sharp, S.B., Villalvazo, M., Espinosa, A., Damle, S., Padilla, X., Hartono, J., Gonzalez, R., and Vu, S. 2002. BC3H1 myogenic cells produce an infectious ecotropic murine leukemia virus. In Vitro Cellular & Developmental Biology-Animal 38: 378-381. |
| Sharp, S.B., Villalvazo, M., Huang, M., Gonzalez, R., Alarcon, I., Bahamonde, M., D'Agostin, D.M., Damle, S., Espinosa, A., Han, S.J., Liu, J., Navarro, P., Salguero, H., Son, J., and Vu, S. 2002. Further characterization of BC3H1 myogenic cells reveals lack of p53 activity and underexpression of several p53 regulated and extracellular matrix-associate gene products. In Vitro Cellular & Developmental Biology-Animal 38: 382-393. |
| Green, N., Vu, S., Farahmand, S., and Sharp, S.B. 1999. Limited T4 exonuclease activity and partial fill-in expand insertion site options for PCR subcloning. Biotechniques 27: 914-916. |
| McQueen, N.L, and Sharp, S.B. 1999. Molecular Diagnostics—an upper division/graduate course. Biochemical Education 27: 145-149. |
Note: ASCL = Wallis Annenberg Integrated Science Complex-Wing A (La Kretz Hall). When calling from off-campus, the area code and prefix for all telephone extensions is (323) 34X-XXXX.